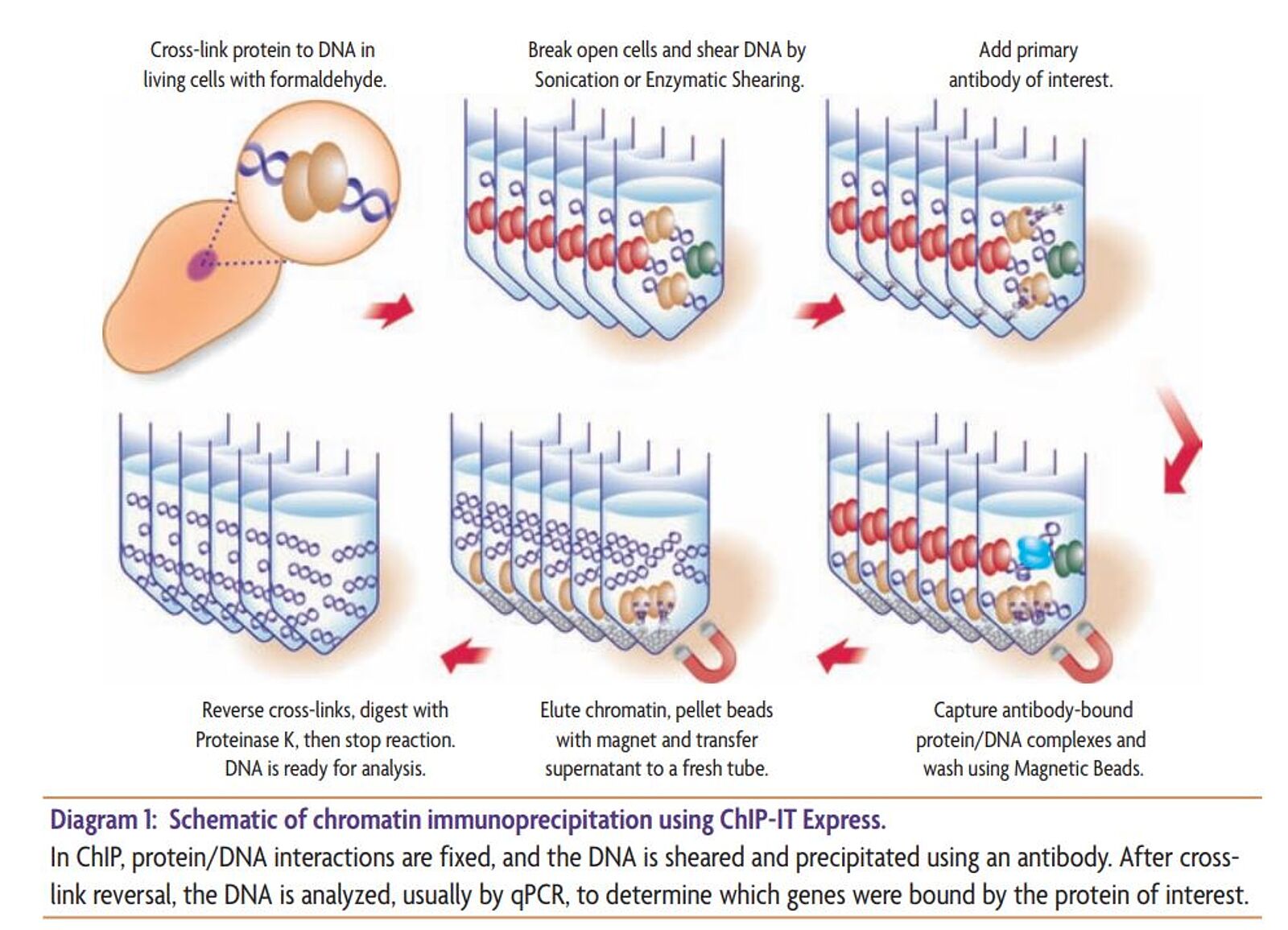
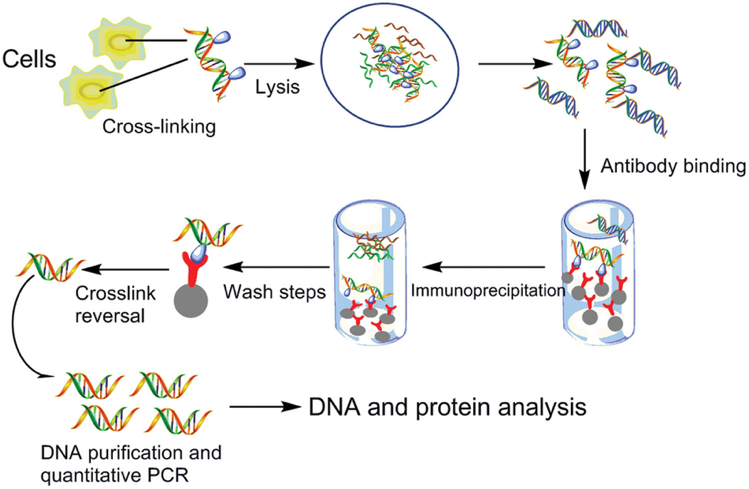
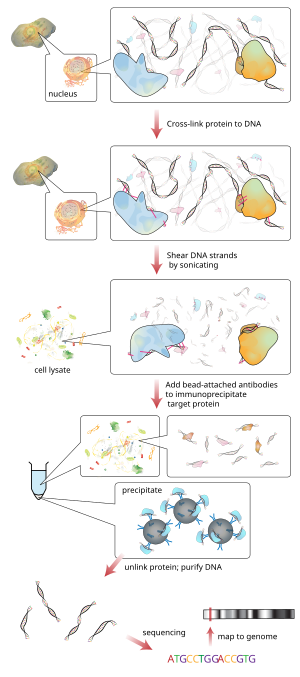
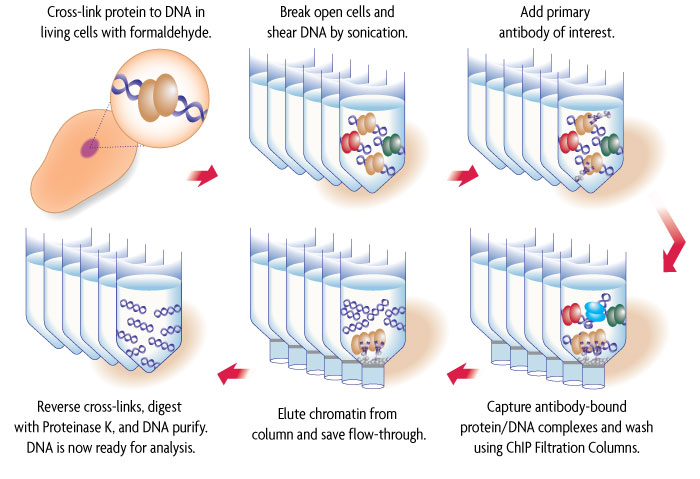
Identification of Putative New Target Genes & Promoter Co-Occupancy Using Novel ChIP Protocols: R&D Systems
The IKAROS Interaction with a Complex Including Chromatin Remodeling and Transcription Elongation Activities Is Required for Hematopoiesis | PLOS Genetics
Enhancer-mediated enrichment of interacting JMJD3–DDX21 to ENPP2 locus prevents R-loop formation and promotes transcription

ChIP and re-ChIP assays. A, top panel, map of the ERV-9 LTR. E, the LTR... | Download Scientific Diagram

Bone-Specific Transcription Factor Runx2 Interacts with the 1α,25-Dihydroxyvitamin D3 Receptor To Up-Regulate Rat Osteocalcin Gene Expression in Osteoblastic Cells | Molecular and Cellular Biology
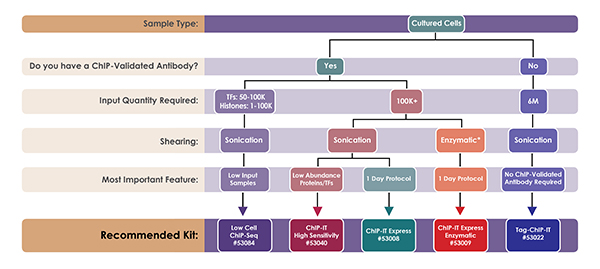
ChIP and Other Methods to Study DNA-Protein Interactions | Biocompare: The Buyer's Guide for Life Scientists

ChIP and Re-ChIP Assays: Investigating Interactions Between Regulatory Proteins, Histone Modifications, and the DNA Sequences to Which They Bind | SpringerLink

Grand Chocolate Chip Cookie | Recipe | Chocolate chip cookies, Ghirardelli recipes, No cook desserts

Identification of Putative New Target Genes & Promoter Co-Occupancy Using Novel ChIP Protocols: R&D Systems