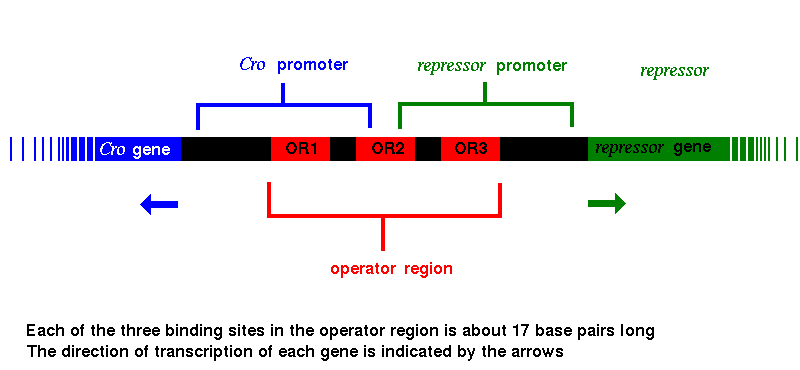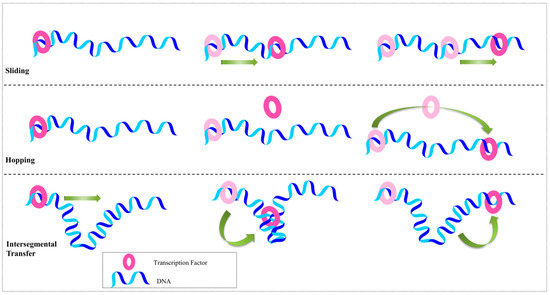
Genes | Free Full-Text | Proteins Recognizing DNA: Structural Uniqueness and Versatility of DNA-Binding Domains in Stem Cell Transcription Factors
A new mode of DNA binding distinguishes Capicua from other HMG-box factors and explains its mutation patterns in cancer | PLOS Genetics
Genome-Wide Profiling of p63 DNA–Binding Sites Identifies an Element that Regulates Gene Expression during Limb Development in the 7q21 SHFM1 Locus | PLOS Genetics

Dna Binding Proteins Transcription Factors - Control Of Gene Expression In Eukaryotes - MCAT Content
Probing the Informational and Regulatory Plasticity of a Transcription Factor DNA–Binding Domain | PLOS Genetics

Cofactor Binding Evokes Latent Differences in DNA Binding Specificity between Hox Proteins - ScienceDirect
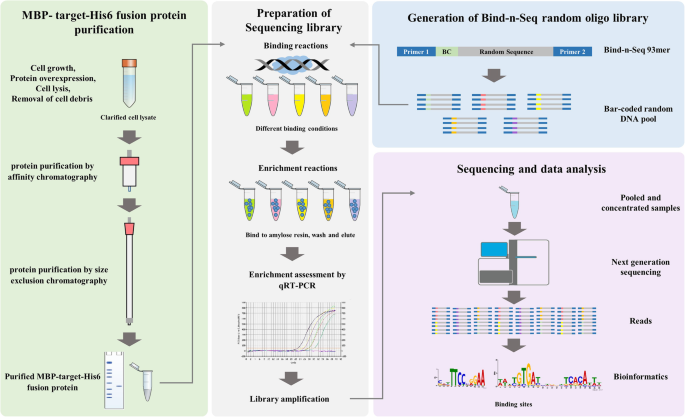
An improved bind-n-seq strategy to determine protein-DNA interactions validated using the bacterial transcriptional regulator YipR | BMC Microbiology | Full Text
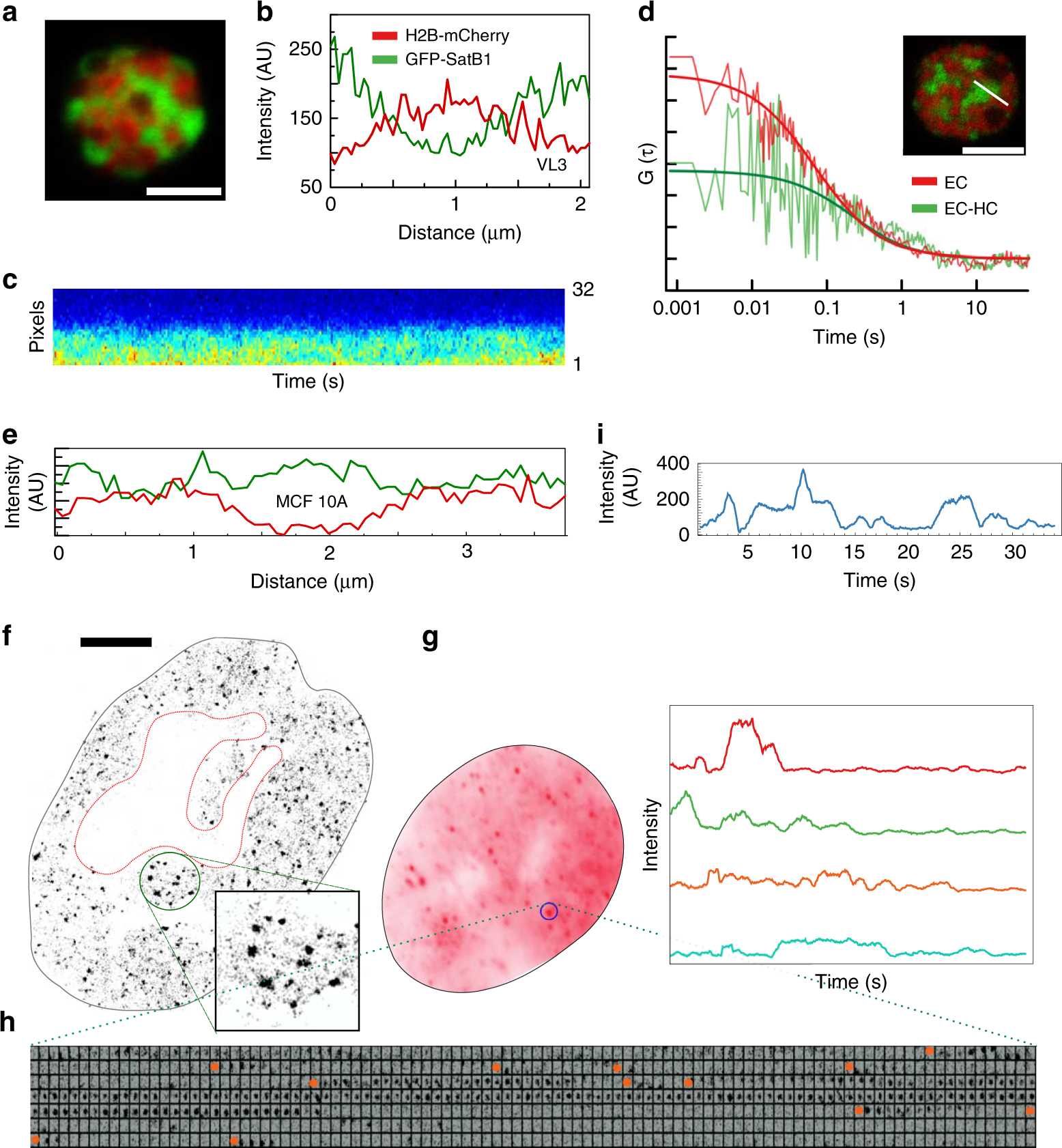
Satb1 integrates DNA binding site geometry and torsional stress to differentially target nucleosome-dense regions | Nature Communications

Coregulation of Transcription Factor Binding and Nucleosome Occupancy through DNA Features of Mammalian Enhancers - ScienceDirect

The Role of Charge Density Coupled DNA Bending in Transcription Factor Sequence Binding Specificity: A Generic Mechanism for Indirect Readout | Journal of the American Chemical Society



![PDF] Protein–DNA binding: complexities and multi-protein codes | Semantic Scholar PDF] Protein–DNA binding: complexities and multi-protein codes | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/da4fc83ebc798bbe5d356dd30cb2ae14fb01ae61/5-Figure1-1.png)